Navigation:
You are here: Home en Product information Introduction (zeige diese Seite in  )
)
Introduction
Dealing with GC/MS chromatograms adds the complexity of comparing the mass spectra of the peaks. Doing this manually further slows down the process of generatiing the results. This makes searching more than a couple of chromatograms impossible.
MSChromSearch can save a tremendous amount of time and offers an unambiguous numerical value for the match between two chromatograms.
MSChromSearch
chromatogram comparison software
MSChromSearch makes a chromatogram library search in a set of other chromatograms, as simple as a library search in a mass spectrum library. The user can select between mass spectrum, retention time, retention index and/or intensity comparisons to be taken into account. From the selected data a “fit value” is calculated, a measure for the similarity between two chromatograms, and chromatograms that were found are listed in the form of a hit list based on the fit value. MSChromSearch enables comparison of a chromatogram with several hundred chromatograms within a very short time.
The first step in using MSChromSearch is analysing the new chromatogram with a given processing method of your data system to get peak and spectra information. The result will be a list of peaks with their chromatographic and spectral data which is usually written in some data system file.
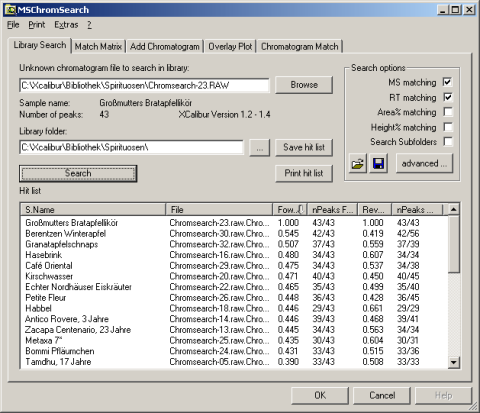
MSChromSearch will read this file and compare these data against the chromatograms in its data base. The result is a fit value which gives the amount of similarity between the two chromatograms. As this process is repeated for each library in the chromatogram database the final step is displaying a hit list sorted by fit value.
This list may be printed or saved as *.csv to open a wider range of post processing options. A simple right click opens the door to further operations like opening unknown and library chromatogram in the QualBrowser of Xcalibur®, or to use deeper analysis within MSChromSearch.
next article:
